Popular Articles
- Earliest molecular events of vision revealed
- Dynamics and Kinetics in Structural Biology
- XFEL Pulses Demonstrate How Plants Perceive Light
- Structural biology is solved -- now what?
- BioXFEL Postdoctoral Fellowship Award
Archived Articles
- Details
- Tuesday, 10 January 2017

A new imaging technique to see proteins live in motion has been developed. The feat achieved by researchers at the UT Southwestern Medical Center comes as a response to electric field pulses and X-rays acting on protein particles.
Given that subtle motions govern protein functions that take place on nano time scales of trillionths of a second, the study offers great potential in knowing how proteins work.
The study's senior author was Rama Ranganathan, Director of the Cecil H. and Ida Green Center for Molecular Biology at UT Southwestern and he is bullish on the outcome.
"So far, we have had no direct way of 'seeing' the motions of amino acids over this range and with atomic precision, which has limited our ability to understand, engineer, and control proteins," Ranganathan said.
The study was done in conjunction with scientists at the University of Chicago and published in Nature.
Use Of EF-X Technology
Using an innovative process called EF-X or Electric field-stimulated X-ray Crystallography, the process allows motions within proteins to be captured visually in real time at atomic resolution and precision.
Creating video-like images of proteins in action has been a milestone, claimed the main author.
This would go a long way in explaining how proteins work in normal and disease states and open more research in drug discovery and protein engineering.
The Real Process
In terms of technique, the process works with X-ray images of protein reactions after applying an electric field. This works on the premise that electromagnetic forces actually govern all protein processes.
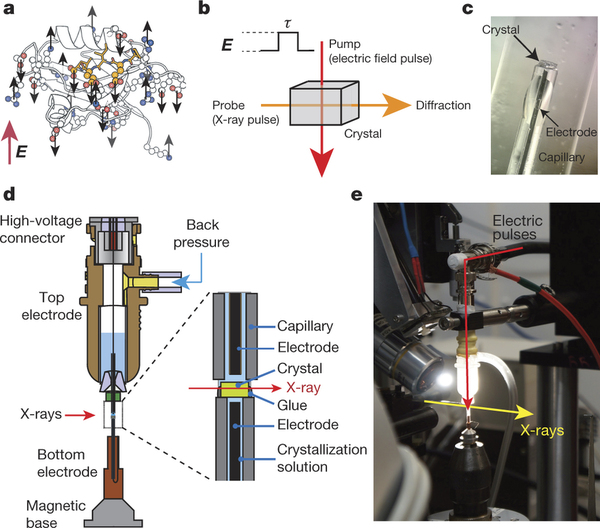
It is done by placing protein crystals between glass capillaries containing a wire followed by application of electric force and X-ray pulses.
Diverse diffraction patterns are then recorded over several intervals including readings before the electric pulse, and again after passing 50, 100 and 200 nanoseconds.
The research team claimed that the technique would work in any protein as a new tool for designing new proteins or drugs.
The technique was claimed to be successful in studying the human ubiquitin ligase protein of the PDZ domain and yielded valuable information on how it works.
Bacterial Protein Boosting Insulin
Meanwhile, a research says insulin-producing beta cells of the pancreas can be stimulated by bacterial protein generated at the zebrafish gut.
This was disclosed by the researchers at the University of Oregon. It may be recalled that Beta cells are endowed with the power of producing insulin for regulating sugar metabolism. The absence of Insulin invites Type 1 diabetes that has afflicted 1.5 million people in the U.S.
The related research was published in the journal eLife.
UO biologist and co-author Karen Guillemin hailed the study and said understanding beta cells and its damage in patients with Type 1 diabetes would support for new preventative and therapeutic approaches.
"We're realizing that the microbiome is a rich source for discovering new biomolecules that have enormous potential for manipulating and promoting our health," said Guillemin.
Original article here





