Popular Articles
- Earliest molecular events of vision revealed
- Dynamics and Kinetics in Structural Biology
- XFEL Pulses Demonstrate How Plants Perceive Light
- Structural biology is solved -- now what?
- BioXFEL Postdoctoral Fellowship Award
Archived Articles
- Details
- Tuesday, 01 August 2017

BioXFEL scientist Vadim Cherezov, along with others, publishes research article in ScienceDirect.
Abstract:
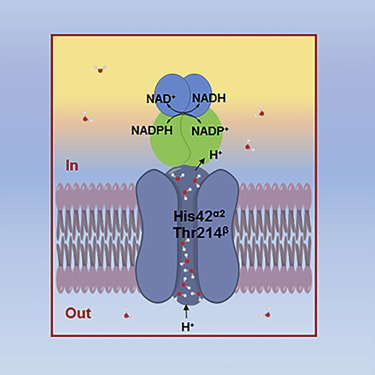
The nicotinamide nucleotide transhydrogenase (TH) is an integral membrane enzyme that uses the proton-motive force to drive hydride transfer from NADH to NADP+ in bacteria and eukaryotes.
Here we solved a 2.2-Å crystal structure of the TH transmembrane domain (Thermus thermophilus) at pH 6.5. This structure exhibits conformational changes of helix positions from a previous structure solved at pH 8.5, and reveals internal water molecules interacting with residues implicated in proton translocation.
Together with molecular dynamics simulations, we show that transient water flows across a narrow pore and a hydrophobic “dry” region in the middle of the membrane channel, with key residues His42α2 (chain A) being protonated and Thr214β(chain B) displaying a conformational change, respectively, to gate the channel access to both cytoplasmic and periplasmic chambers. Mutation of Thr214β to Ala deactivated the enzyme. These data provide new insights into the gating mechanism of proton translocation in TH.





