LCLS serial femtosecond crystallography data analysis instructions
Data Analysis
This resource belongs to the Data Analysis group.
Category
Published on
Abstract
The links below will take you to detailed instructions and sound advice from experienced LCLS data analysts. These pages are focused on serial femtosecond crystallography (SFX), but much of it applies to other X-ray diffraction or imaging experiments at LCLS.
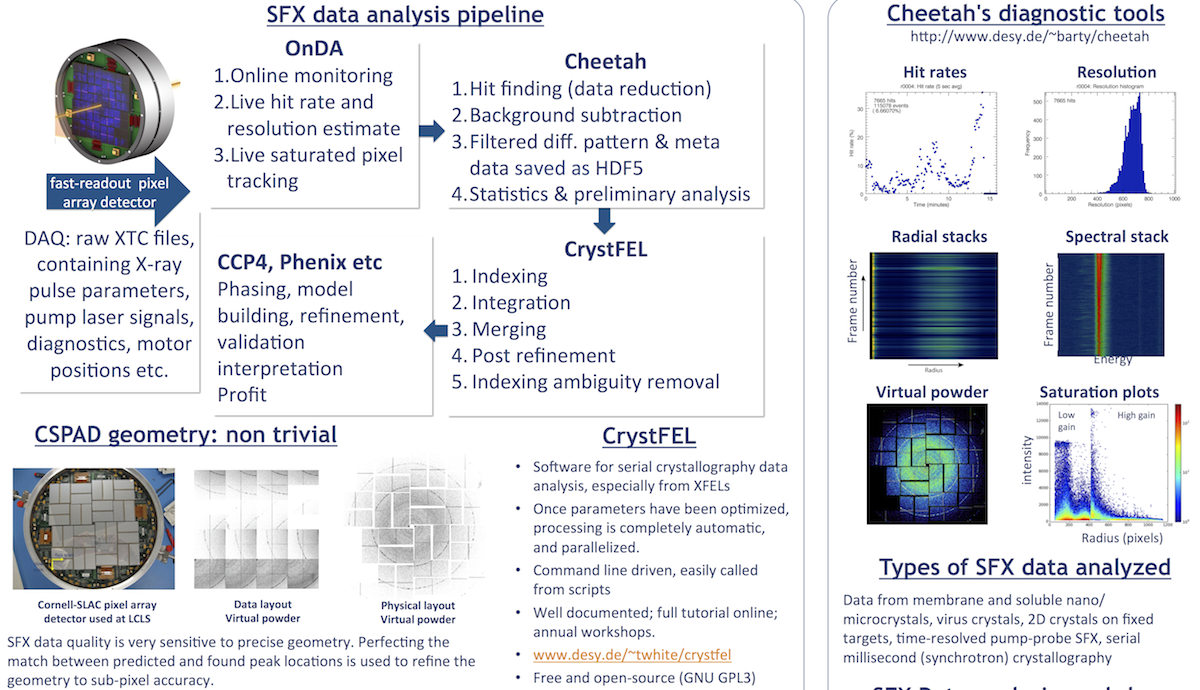
- Brief introduction to serial femtosecond crystallography & slides from "Software for serial femtosecond crystallography data analysis" from IUCr 2014.
- Before your beamtime
- During your beamtime
- Accessing your data at LCLS
- Detectors and geometry
- Data reduction and hit finding: Cheetah
- Live monitoring and hit finding parameter tweaker: OnDA
- Working with LCLS data remotely
- Managing your LCLS data after the beamtime
- Indexing, merging, evaluating crystal diffraction data: CrystFEL
The SFX data analysis pipeline we will be referring to is outlined here. These pages will describe the CSPAD/DAQ, Cheetah and CrystFEL steps.
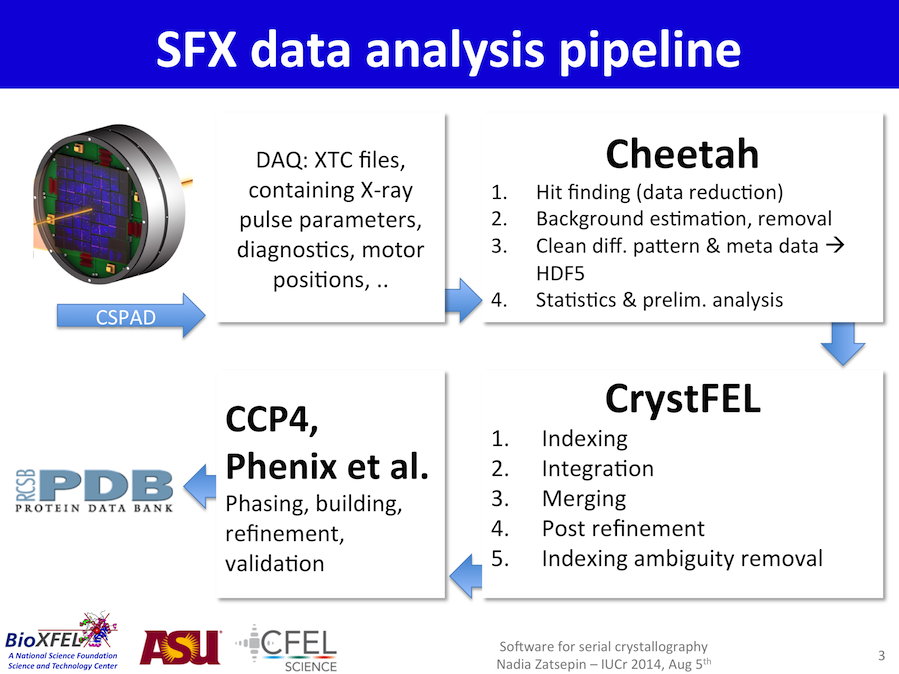
Additionally, you will find links to
- Useful data analysis scripts, all free for your use and modification. These were contributed by BioXFEL members and collaborators.
- 2015 Data Analysis Workshop at ACA, Philadelphia, including PDF's of presentations.
- 2015 Phenix workshop at Rice University, to help with analyzing your merged serial crystallography data after CrystFEL
- Videos and presentations from our 2014 data analysis (serial femtosecond crystallography) workshop held at LBNL
- Presentation by BioXFEL researcher Marc Messerschmidt about crystallography with an XFEL.
- Cheetah documentation: page 1, page 2, keywords and hit finding algorithms.
If you see typos or errors or would like more detail about a particular topic, please email the authors (top of this page).
References
1. LCLS Data Analysis pages (getting started, python modules, detector calibration GUI, TimeTool etc): https://confluence.slac.stanford.edu/display/PSDM/LCLS+Data+Analysis 2. SFX Data analysis workshop at ACA 2015 Workshop spreadsheet 3. BioXFEL data analysis group : https://www.bioxfel.org/groups/dataanalysis 4. Resources for BioXFEL data analysis group: https://www.bioxfel.org/groups/dataanalysis/resources 5. GitHub repository of scripts: https://github.com/bioxfel/scripts




